Genetics, Bioinformatics and Systems Biology Colloquium
Thursdays, 12:00 pm – 1:00 pm
UC San Diego, Powell-Focht Bioengineering Hall, Fung Auditorium
Complete schedule here
Friday, April 17, 2015 from 8 am – 5 pm
Main Auditorium at the Sanford Consortium for Regenerative Medicine
Organized by Barry Demchak and Jason Kreisberg
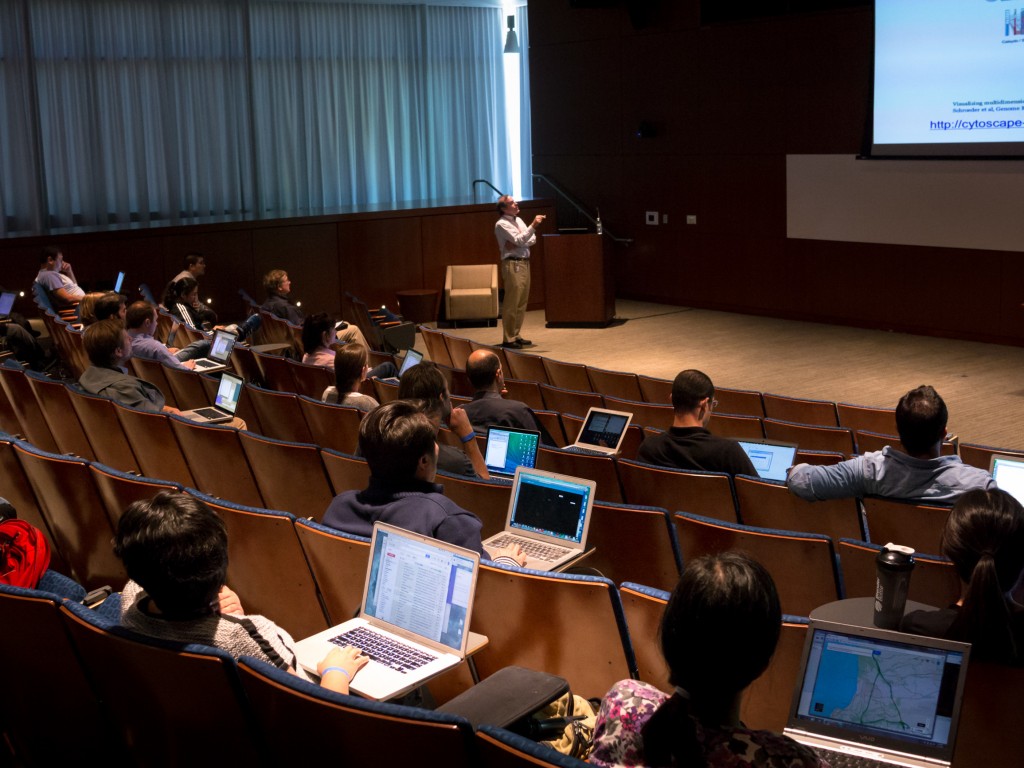
Network analysis has become an increasingly common approach for tackling complicated, large data sets. In the morning, this workshop will feature a number of speakers discussing their efforts to use network analysis to address pressing biological questions and unmet medical needs. One frequently used tool for network analysis is Cytoscape, an open source bioinformatics software platform for visualizing molecular interaction networks and biological pathways and for integrating these networks with annotations, gene expression profiles and other state data. Although Cytoscape was originally designed for biological research, now it is a general platform for complex network analysis and visualization. In the afternoon, the Cytoscape development team from Trey Ideker’s lab will present a series of demonstrations and workshops covering a range of issues for both novice and advanced users.
Additional information on Cytoscape can be found on the Cytoscape home page, the Cytoscape Wiki or Wikipedia.
Please visit here to register.
New this year
Cytoscape clinic
Lunchtime roundtable discussion on the emerging Cytoscape Cyberinfrastructure, which leverages Cytoscape and web-based services to enable advanced workflows and promote reproducibility.
New in Cytoscape this year
New publishing models for web – viewable networks
Nodes drawn as charts – a new dimension in visualization
Powerful REST interface enables liaison with Python, R and others
… and more!
Schedule
| 8:00 | Registration |
| 9:00 | Barry Demchak (Chief Architect, Cytoscape) and Jason Kreisberg (Assistant Director, SDCSB) Welcome |
| 9:10 | Nik Schork (Director of Human Biology, JCVI) Rewiring genetic networks and drug targeting |
| 9:50 | Rob Quinn (Post-Doctoral Fellow, Forest Rohwer’s Lab, SDSU) Network approaches link microbial and host physiology in the cystic fibrosis lung |
| 10:10 | Lars Pache (Post-Doctoral Fellow, Sumit Chanda’s Lab, SBP) Network-based analysis of host-pathogen interactions |
| 10:30 | Coffee |
| 10:50 | Rommie Amaro (Associate Professor, Dept of Chem & Biochem, UCSD) Allosteric networks through the lens of a computational microscope |
| 11:30 | S. Carol Huang (Post-Doctoral Fellow, Joe Ecker’s Lab, Salk) Multiscale network analysis for biological insights in humans and plants |
| 11:50 | Keiichiro Ono (Cytoscape Core Developer, Trey Ideker’s Lab, UCSD) What’s new in cytoscape |
| 12:30 | Lunch |
| 12:30 | Barry Demchak, Keiichiro Ono and Dexter Pratt (Director of the NDEx Project, Trey Ideker’s Lab, UCSD) Cytoscape CyberInfrastructure Roundtable (Lobby Conference Room) |
| 1:30 | Barry Demchak, Rintaro Saito (Project Scientist, Kumar Sharma’s Lab, UCSD) and Tim Hull (Cytoscape Developer, Trey Ideker’s Lab, UCSD) Basic Cytoscape (Auditorium) |
| 1:30 | Keiichiro Ono Advanced Cytoscape ** (Lobby Conference Room) |
| 3:00 | Barry Demchak, Keiichiro Ono, Dexter Pratt, Rintaro Saito and Tim Hull Cytoscape Clinic (Lobby Conference Room) |
Key Notes
Participants should bring their own laptop with an installed copy of the latest Cytoscape (v3.2.1; the workshop will cover installation for anyone having difficulties), as well as your own data to experiment with. Note that we’ll offer hands-on help during the Cytoscape Clinic. Finally, please see suggested reading before the workshop. If you have questions, please contact Barry Demchak.
There will be only limited WiFi available during the workshop, and it may be insufficient to support Cytoscape installation during the workshop. PLEASE install the latest Cytoscape ahead of time.
** Advanced Cytoscape ** tutorial will focus on the reproducibility of your data visualization workflows with Cytoscape. This means you will learn how to use Cytoscape with other popular tools such as IPython Notebook or NetworkX. The session will be a shorter version of this 3.5-hour-long tutorial found here. If you are not sure whether you should attend this or not, please send your questions to Keiichiro Ono.
No Show Policy
While we do not charge a fee to attend this event, these programs would not be sustainable and available to all wanting to attend unless all registrants abide by the 48 hour cancellation notice policy. A no-show can limit your ability to book our events in the future. Thank you in advance for your cooperation.
Directions
Free parking at the Sanford Consortium will be provided for registered attendees. There is also additional free parking near the Torrey Pines Gliderport, which is within walking distance. The UCSD Shuttle Route has two drop off locations within walking distance to the Sanford Consortium.
Pictures of scientists having fun
With additional support provided by Flu-OMICS
Flu-OMICS is funded by a grant from the NIH/NIAID U19AI106754