Genetics, Bioinformatics and Systems Biology Colloquium
Thursdays, 12:00 pm – 1:00 pm
UC San Diego, Powell-Focht Bioengineering Hall, Fung Auditorium
Complete schedule here
The International Genetically Engineered Machines (iGEM) Competition is the premiere student competition in Synthetic Biology. Teams from top universities around the world compete solve problems in synthetic biology using a kit of biological parts. Past projects have included producing a rainbow of pigmented bacteria, banana and wintergreen scented bacteria, bacteria capable of sensing arsenic in drinking water. In recent years, new tracks have been added to competition including one for computational biology.
2015
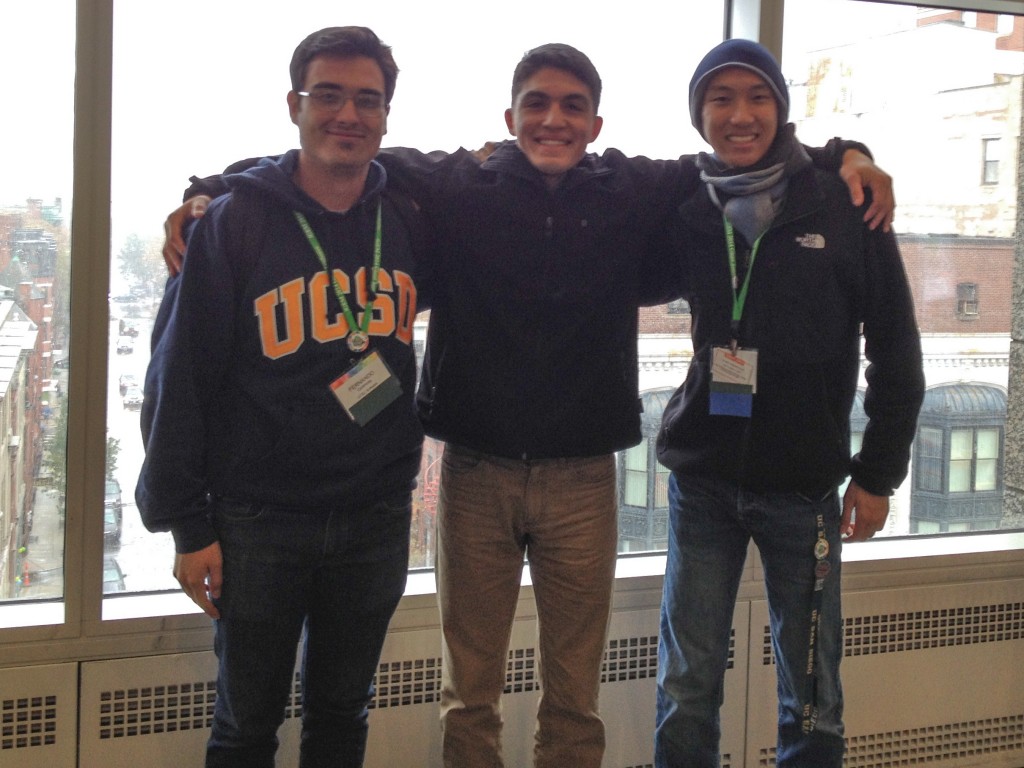
UC San Diego’s 2015 iGEM team is moving in an exciting new direction with the incorporation of a wet lab component in this summer’s project. Supported by mentors from the Neural Interaction Laboratory, Systems Biology Research Group and the BioCircuits Institute, the team’s six undergraduates will be working to optimize metabolic pathways through the rapid identification of rate-limiting steps. To achieve this goal, they will conduct experiments and build mathematical models of the metabolic pathways. In late September, the team will be representing UCSD on an international level and presenting their completed work in Boston at the 11th annual iGEM competition.
Because it exposes undergraduates to the type of work that they will likely pursue in their future, iGEM also presents a powerful educational tool for future scientists and engineers. This year’s team is composed of members of all skill levels with a wide range of backgrounds including bioengineering, biochemistry, cell biology and physics. They are working together intensely and developing the skills needed to work in an interdisciplinary environment. A wiki detailing their progress can be found here.
2014
UCSD’s computational iGEM team in 2014 consisted of 10 undergraduates with mentoring was provided by 4 graduate student. The students spent the summer building SBiDer, a modern web application that leverages a novel database that captures the interaction between genetic circuits to build complex genetic circuits. A demo video is available here and a wiki detailing the project design is here. In the fall, the students presented their work at the iGEM Jamboree in Boston where they won a Gold Medal, indicating a high level of performance on par with other top universities such as MIT, Cornell and Stanford. After the Jamboree, the undergraduates also participated in the second Network of BioThings Hackathon, placing second amongst a field of teams composed primarily of professional developers, graduate students, postdoctoral fellows and faculty members.
Perhaps most importantly, the students learned new skills and developed a genuine interest in synthetic biology and appreciation for interdisciplinary research. Most of the students that programmed the web application had no programming background; the computationally oriented students were able to pick up synthetic biology concepts. Six of the 10 students have indicated an inclination to pursue graduate studies in the future.
The undergraduates are currently finishing the software and building compatibility with NDex, a online repository of network data. Once the software is stable, it will be deployed on a server provided by the Ideker lab. We are hopeful that iGEM HQ or COSBI (a synthetic biology consortium) would be interested in continuing development on our web application.